Article of the week: A four‐group urine risk classifier for predicting outcomes in patients with prostate cancer
Every week, the Editor-in-Chief selects an Article of the Week from the current issue of BJUI. The abstract is reproduced below and you can click on the button to read the full article, which is freely available to all readers for at least 30 days from the time of this post.
In addition to the article itself, there is an editorial written by a prominent member of the urological community. These are intended to provoke comment and discussion and we invite you to use the comment tools at the bottom of each post to join the conversation.
If you only have time to read one article this week, it should be this one.
A four‐group urine risk classifier for predicting outcomes in patients with prostate cancer
Shea P. Connell*, Marcelino Yazbek-Hanna*, Frank McCarthy†, Rachel Hurst*, MartynWebb*, Helen Curley*, Helen Walker‡, Rob Mills‡, Richard Y. Ball‡, Martin G. Sanda§, Kathryn L. Pellegrini§, Dattatraya Patil§, Antoinette S. Perry¶, Jack Schalken**, Hardev Pandha††, Hayley Whitaker‡‡, Nening Dennis†, Christine Stuttle†, Ian G. Mills§§¶¶***, Ingrid Guldvik¶¶, Movember GAP1 Urine Biomarker Consortium1, Chris Parker†††, Daniel S. Brewer*‡‡‡, Colin S. Cooper* and Jeremy Clark*
*Norwich Medical School, University of East Anglia, Norwich, †Institute of Cancer Research, Sutton, ‡Norfolk and Norwich University Hospitals NHS Foundation Trust, Norwich, UK, §Department of Urology, Winship Cancer Institute, Emory University School of Medicine, Atlanta, GA, USA , ¶School of Biology and Environmental Science, Science West, University College Dublin, Dublin 4, Ireland, **Nijmegen Medical Centre, Radboud University Medical Centre, Nijmegen, The Netherlands, ††Faculty of Health and Medical Sciences, The University of Surrey, Guildford, ‡‡Molecular Diagnostics and Therapeutics Group, University College London, London, §§School of Medicine, Dentistry and Biomedical Sciences, Institute for Health Sciences, Centre for Cancer Research and Cell Biology, Queen’s University Belfast, Belfast, UK, ¶¶Centre for Molecular Medicine, University of Oslo, Oslo, Norway, ***Nuffield Department of Surgical Sciences, University of Oxford, Oxford, †††Royal Marsden Hospital, Sutton and ‡‡‡Earlham Institute, Norwich, UK
Abstract
Objectives
To develop a risk classifier using urine‐derived extracellular vesicle (EV)‐RNA capable of providing diagnostic information on disease status prior to biopsy, and prognostic information for men on active surveillance (AS).
Patients and Methods
Post‐digital rectal examination urine‐derived EV‐RNA expression profiles (n = 535, multiple centres) were interrogated with a curated NanoString panel. A LASSO‐based continuation ratio model was built to generate four prostate urine risk (PUR) signatures for predicting the probability of normal tissue (PUR‐1), D’Amico low‐risk (PUR‐2), intermediate‐risk (PUR‐3), and high‐risk (PUR‐4) prostate cancer. This model was applied to a test cohort (n = 177) for diagnostic evaluation, and to an AS sub‐cohort (n = 87) for prognostic evaluation.
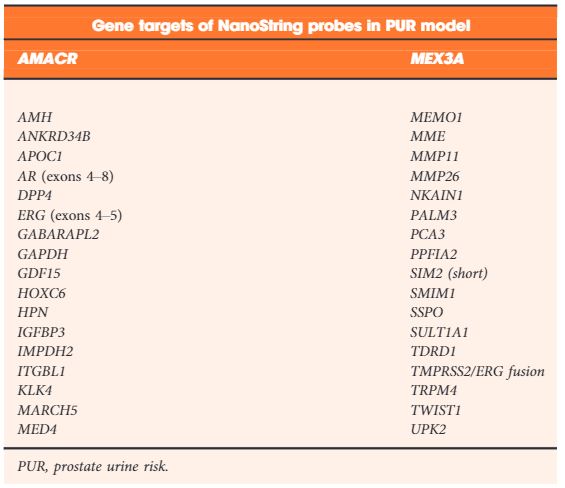
Table 2. NanoString gene probes incorporated by LASSO regularization in the final optimal model used to produce the prostate urine risk signatures
Results
Each PUR signature was significantly associated with its corresponding clinical category (P < 0.001). PUR‐4 status predicted the presence of clinically significant intermediate‐ or high‐risk disease (area under the curve = 0.77, 95% confidence interval [CI] 0.70–0.84). Application of PUR provided a net benefit over current clinical practice. In an AS sub‐cohort (n = 87), groups defined by PUR status and proportion of PUR‐4 had a significant association with time to progression (interquartile range hazard ratio [HR] 2.86, 95% CI 1.83–4.47; P < 0.001). PUR‐4, when used continuously, dichotomized patient groups with differential progression rates of 10% and 60% 5 years after urine collection (HR 8.23, 95% CI 3.26–20.81; P < 0.001).
Conclusion
Urine‐derived EV‐RNA can provide diagnostic information on aggressive prostate cancer prior to biopsy, and prognostic information for men on AS. PUR represents a new and versatile biomarker that could result in substantial alterations to current treatment of patients with prostate cancer.