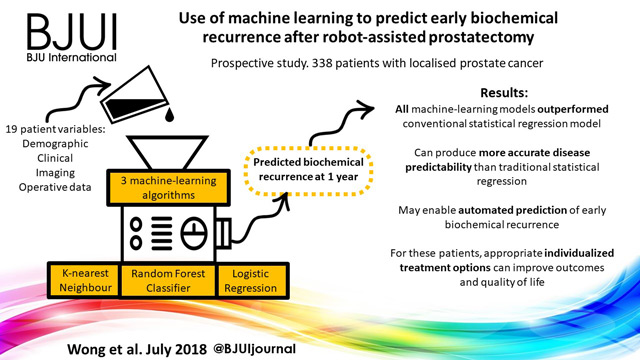
Article of the Month: Use of machine learning to predict early biochemical recurrence after robot‐assisted prostatectomy
Every month, the Editor-in-Chief selects an Article of the Month from the current issue of BJUI. The abstract is reproduced below and you can click on the button to read the full article, which is freely available to all readers for at least 30 days from the time of this post.
In addition to the article itself, there is an accompanying editorial written by a prominent member of the urological community. This blog is intended to provoke comment and discussion and we invite you to use the comment tools at the bottom of each post to join the conversation.
If you only have time to read one article this week, it should be this one.
Use of machine learning to predict early biochemical recurrence after robot‐assisted prostatectomy
Nathan C. Wong , Cameron Lam, Lisa Patterson and Bobby Shayegan
Division of Urology, Department of Surgery, McMaster University, Hamilton, ON, Canada
Abstract
Objectives
To train and compare machine‐learning algorithms with traditional regression analysis for the prediction of early biochemical recurrence after robot‐assisted prostatectomy.
Patients and Methods
A prospectively collected dataset of 338 patients who underwent robot‐assisted prostatectomy for localized prostate cancer was examined. We used three supervised machine‐learning algorithms and 19 different training variables (demographic, clinical, imaging and operative data) in a hypothesis‐free manner to build models that could predict patients with biochemical recurrence at 1 year. We also performed traditional Cox regression analysis for comparison.
Results
K‐nearest neighbour, logistic regression and random forest classifier were used as machine‐learning models. Classic Cox regression analysis had an area under the curve (AUC) of 0.865 for the prediction of biochemical recurrence. All three of our machine‐learning models (K‐nearest neighbour (AUC 0.903), random forest tree (AUC 0.924) and logistic regression (AUC 0.940) outperformed the conventional statistical regression model. Accuracy prediction scores for K‐nearest neighbour, random forest tree and logistic regression were 0.976, 0.953 and 0.976, respectively.
Conclusions
Machine‐learning techniques can produce accurate disease predictability better that traditional statistical regression. These tools may prove clinically useful for the automated prediction of patients who develop early biochemical recurrence after robot‐assisted prostatectomy. For these patients, appropriate individualized treatment options can improve outcomes and quality of life.