Editorial: Is transrectal ultrasonography of the prostate obsolete in the MRI era?
Sampling of prostate tissue to confirm pathologically a clinical suspicion of cancer has undergone an exponential change. The random systematic prostate biopsy technique was the only method used for many decades, initially guided by the finger but, since 1989, performed with TRUS guidance. Now, within the space of only a few years, we have entered the era of performing prostate biopsies on the basis of high‐tech three‐dimensional multiparametric MRI images, including software that can track the exact course of the biopsy needle [1]. While new technical developments in general lead to better, more individually directed healthcare, there is always the risk of abandoning ‘old’ but well developed and extensively tested techniques too soon. In this issue of the BJUI, Press et al. [2] looked at the added value of the presence of an ‘old‐fashioned’ TRUS‐detected lesion in cancer‐suspicious regions on MRI to better predict the presence of clinically significant prostate cancer (csPCa) defined as Gleason score ≥7. In their study comprising 1058 men, it was shown that a well‐demarcated abnormal TRUS finding noted at the time of MRI‐TRUS fusion‐guided prostate biopsy coincides with an increased risk of csPCa detection, independent of MRI suspicion (Prostate Imaging Reporting and Data System [PI‐RADS] score).
Increasing PI‐RADS score is correlated with an increased percentage of csPCa after targeted biopsy, both at initial and repeat biopsy. In a review based on data from 8252 men, it was shown that there is a gradual increase in the detection of csPCa from PI‐RADS 3 to PI‐RADS 4 to PI‐RADS 5 index lesions. For example, at first biopsy, the overall rate of PCa detection and the percentage of csPCa were 39%, 62% and 92% and 54%, 63% and 76% for PI‐RADS 3, 4 and 5 lesions, respectively. This means that in men with PI‐RADS 3 lesions, representing approximately one‐third of men deemed eligible for further assessment, only 39% will be diagnosed with PCa and half of the PCa detected will be potentially indolent Gleason 6 PCa [3]. This makes this group of men extremely interesting for further risk stratification before biopsy. Multivariable risk stratification in which PSA density plays an important role has been shown to be of value in these men [4] but further refinement could potentially be made by including suspicious lesions identified at TRUS.
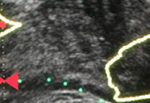
Apart from the added value of TRUS findings in terms of risk stratification, the performance of the MRI‐targeted biopsy itself could be improved by visual guidance of hypoechoic lesions. In the present study by Press et al [2], a hypoechoic TRUS lesion was present at or near the location of two‐thirds of cancer‐suspicious lesions on MRI. The authors more or less advise to direct the targeted biopsy cores not only to the MRI suspicious lesion, but also the TRUS suspicious lesion, both of which often do not fully overlay in a software‐assisted MRI‐TRUS fusion model. The extent to which this ‘correction for misregistration’ is already included during targeted biopsy in current clinical practice is unknown. Although feasible and seemingly important during software‐assisted fusion targeted biopsy, TRUS lesions in cancer‐suspicious MRI regions might be more frequently targeted during cognitive fusion‐targeted biopsy. Two recent studies underline the important message of the present study, and show that a considerable proportion of csPCa is missed in and around MRI‐suspicious lesions by targeted biopsies, as a result of sampling errors related to both misregistration and intra‐tumour heterogeneity [5, 6]. As suggested by these studies, visual guidance by hypoechoic lesions and ‘focal saturation’ biopsy by additional (peri‐)lesional cores might improve the detection of csPCa.
In summary, ‘good old’ TRUS could be of value in those patients who are virtually always present in scenarios in which a grading system is being used, i.e. patients belonging to the so‐called grey zone. The challenge of risk stratification (i.e. personalized medicine) is to nibble at both sides of the grey zone by implementing new techniques or, more likely by implementing a combination of all available and relevant knowledge.
by Monique J. Roobol, Frank-Jan H. Drost and Arnout R. Alberts
References
- , , et al. The current state of MR imaging‐targeted biopsy techniques for detection of prostate cancer. Radiology 2017; 285: 343–56
- , , , . The ultrasound characteristics of MRI suspicious regions predict the likelihood of clinically significant cancer on MRI‐ultrasound fusion targeted biopsy. BJUI 2019; 123: 439–46.
- . MRI in early prostate cancer detection: how to manage indeterminate or equivocal PI‐RADS 3 lesions? Transl Androl Urol 2018; 7: 70–82
- , , , , , . Risk‐based patient selection for magnetic resonance imaging‐targeted prostate biopsy after negative transrectal ultrasound‐guided random biopsy avoids unnecessary magnetic resonance imaging scans. Eur Urol 2016; 69: 1129–34
- , , et al. Accuracy of transperineal targeted prostate biopsies, visual estimation and image fusion in men needing repeat biopsy in the PICTURE trial. J Urol 2018; 200: 1227–34
- Leest, M, , et al. Head‐to‐head comparison of transrectal ultrasound‐guided prostate biopsy versus multiparametric prostate resonance imaging with subsequent magnetic resonance‐guided biopsy in biopsy‐naive men with elevated prostate‐specific antigen: a large prospective multicenter clinical study. Eur Urol 2018; [Epub ahead of print]. https://doi.org/10.1016/j.eururo.2018.11.023.