Editorial: The utility of microRNAs as biomarkers in predicting progression and survival in patients with clear-cell renal cell carcinoma
RCC constitutes a diverse group of malignancies, yet the clear-cell subtype comprises ~80% of all diagnosed RCC cases [1]. The widespread use of abdominal imaging and subsequent stage migration has resulted in improved RCC 5-year cancer-specific survival. However, the overall mortality of RCC remains largely unchanged [2] and one-third of the patients have metastatic disease at the time of presentation [3]. Accordingly, the ability to precisely predict patient outcome has become an increasingly significant question in the management of these patients with RCC.
Accruing evidence suggests that changes in various biomarkers and their consequent downstream pathways affect cancer initiation and progression. Therefore, accurate prediction of the outcome and prognosis after treatment is necessary [4]. MicroRNAs (miRNAs) are small non-coding RNA molecules that can have significant functions in tumorigenesis [5]. Because of their ability in post-transcriptional regulation of gene expression, tumour-specific genetic defects in miRNA biogenesis and production correlate with development of human cancers. Thus, the differential expression of specific miRNA signatures in different tumours might become an important tool to help in directing cancer diagnosis and treatment [5].
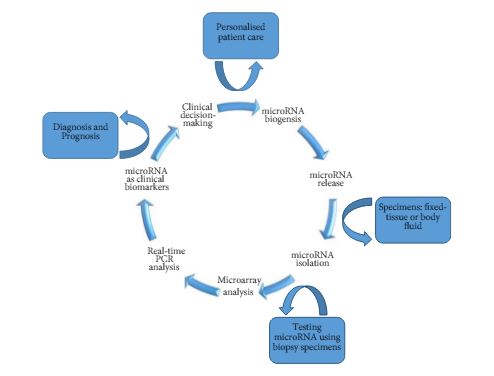
As such, Kowalik et al. [6] report on profiling miRNA to identify biomarker signatures predictive of clear-cell RCC (ccRCC) progression and survival. The authors used 202 formalin-fixed paraffin-embedded samples to isolate RNA from nephrectomy and biopsy specimens (n = 156 and n = 46, respectively) (Fig. 1).
Figure 1. Schematic diagram of miRNA-based biomarkers and potential utility in clinical decision-making approach for targeted therapy and RCC personalised treatment.
The primary analysis of their study [6] focused on the identification of miRNA signatures capable of differentiating between benign and ccRCC, as well as discerning those patients with a non-progressive ccRCC from a progressive clear-cell subtype. The secondary outcome examined the association of miRNA profiles discovered on cancer-specific survival.
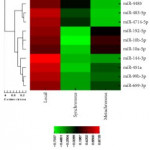
In their initial microarray screening 20 differentially expressed miRNAs, comparing non-progressive with progressive tumours, were identified. The authors found four miRNA panels (10a-5p, 10b-5p, 106a-5p, and 142-5p) as a potential biomarker signature. This model was validated in nephrectomy specimens and resulted in a sensitivity of 86.7%, a specificity of 92.9%, and an area under the curve (AUC) of 0.930 for detecting ccRCC. Further analysis revealed a second signature of two biomarkers (miR-10a-5p and -223-3p) with 93.8% sensitivity, 83.3% specificity, and an AUC of 0.932 when validated for detecting progressive ccRCC. Similarly, the differential expression of these biomarkers could delineate cancer status in biopsy specimens. For correlation of miRNA expression levels with cancer-specific survival, higher expression levels of (miR-10a-5p and miR-10b-5p) and a lower expression level of (miR-223-3p) were significantly associated with survival (P< 0.001), and the median survival times were not reached.
In conclusion, the lack of precise prediction tools has led the authors to explore the potential utility of miRNAs as biomarkers to detect disease presence, biological aggressiveness, and prognosis in ccRCC. However, until future multicentre large prospective studies validate the results of the present work, the transition of miRNA from bench to bedside is emerging on the horizon and has encouraged urologists and scientists to pursue intense translational research in the field. The ability to use miRNAs as biomarkers might be promising for diagnostic and prognostic purposes. These biomarkers may exemplify different aspects of RCC pathogenesis and may potentially have important therapeutic implications to help in a clinical decision-making approach for targeted therapy and RCC personalised treatment.