Article of the Week: Evaluation of targeted and systematic biopsies using MRI and US image-fusion guided transperineal prostate biopsy
Every Week the Editor-in-Chief selects an Article of the Week from the current issue of BJUI. The abstract is reproduced below and you can click on the button to read the full article, which is freely available to all readers for at least 30 days from the time of this post.
In addition to the article itself, there is an accompanying editorial written by a prominent member of the urological community. This blog is intended to provoke comment and discussion and we invite you to use the comment tools at the bottom of each post to join the conversation.
If you only have time to read one article this week, it should be this one.
Multicentre evaluation of targeted and systematic biopsies using magnetic resonance and ultrasound image-fusion guided transperineal prostate biopsy in patients with a previous negative biopsy
Abstract
Objectives
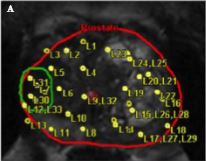
To evaluate the detection rates of targeted and systematic biopsies in magnetic resonance imaging (MRI) and ultrasound (US) image-fusion transperineal prostate biopsy for patients with previous benign transrectal biopsies in two high-volume centres.
Patients and Methods
A two centre prospective outcome study of 487 patients with previous benign biopsies that underwent transperineal MRI/US fusion-guided targeted and systematic saturation biopsy from 2012 to 2015. Multiparametric MRI (mpMRI) was reported according to Prostate Imaging Reporting and Data System (PI-RADS) Version 1. Detection of Gleason score 7–10 prostate cancer on biopsy was the primary outcome. Positive (PPV) and negative (NPV) predictive values including 95% confidence intervals (95% CIs) were calculated. Detection rates of targeted and systematic biopsies were compared using McNemar’s test.
Results
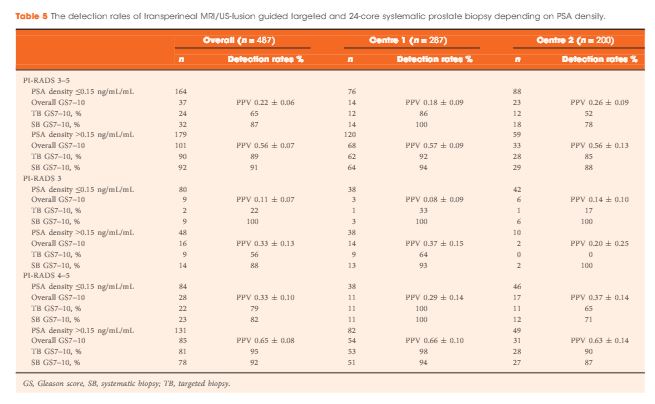
The median (interquartile range) PSA level was 9.0 (6.7–13.4) ng/mL. PI-RADS 3–5 mpMRI lesions were reported in 343 (70%) patients and Gleason score 7–10 prostate cancer was detected in 149 (31%). The PPV (95% CI) for detecting Gleason score 7–10 prostate cancer was 0.20 (±0.07) for PI-RADS 3, 0.32 (±0.09) for PI-RADS 4, and 0.70 (±0.08) for PI-RADS 5. The NPV (95% CI) of PI-RADS 1–2 was 0.92 (±0.04) for Gleason score 7–10 and 0.99 (±0.02) for Gleason score ≥4 + 3 cancer. Systematic biopsies alone found 125/138 (91%) Gleason score 7–10 cancers. In patients with suspicious lesions (PI-RADS 4–5) on mpMRI, systematic biopsies would not have detected 12/113 significant prostate cancers (11%), while targeted biopsies alone would have failed to diagnose 10/113 (9%). In equivocal lesions (PI-RADS 3), targeted biopsy alone would not have diagnosed 14/25 (56%) of Gleason score 7–10 cancers, whereas systematic biopsies alone would have missed 1/25 (4%). Combination with PSA density improved the area under the curve of PI-RADS from 0.822 to 0.846.
Conclusion
In patients with high probability mpMRI lesions, the highest detection rates of Gleason score 7–10 cancer still required combined targeted and systematic MRI/US image-fusion; however, systematic biopsy alone may be sufficient in patients with equivocal lesions. Repeated prostate biopsies may not be needed at all for patients with a low PSA density and a negative mpMRI read by experienced radiologists.